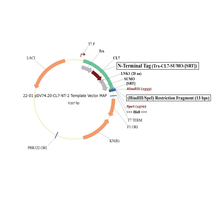
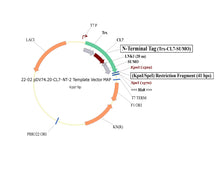
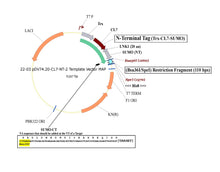
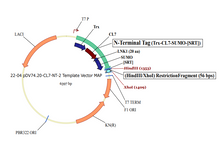
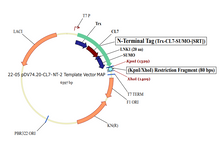
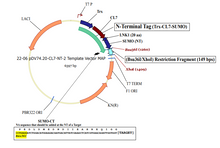
Plasmid #22 enables the tagging of target proteins with a cleavable CL7 at the N-terminus and an uncleavable His8 at the C-terminus. There are three options for the 5’ placement of target genes (see scheme below), and SpeI is the 3’ cloning site.
SKU: 20-1022
EXPRESSION
Transcription is induced with IPTG and driven by the T7 RNA polymerase. The plasmid is designed for expression in E. coli.
PEPTIDE TAG
The N-terminus of the protein is tagged with CL7.
CLEAVAGE SITE(S)
SUMO and Sortase A cleavage sites are both located between the CL7 tag and the target protein’s N-terminus.
OTHER TAGS
The coding region begins with a Trx tag and ends with a His8 tag in Plasmid #22.
CLONING OPTIONS
1. HindIII/SpeI Insertion Site – Trx | CL7 | 20-Amino Acid Linker | SUMO | SRT | 13-Amino Acid Linker | Gene of Interest | His8 (This is the same as option 4, with the exception of His8)
2. KpnI/SpeI Insertion Site – Trx | CL7 | 20-Amino Acid Linker | SUMO | 3| Gene of Interest | His8 (This is the same as option 5, with the exception of His8)
3. Bsu36I/SpeI Insertion Site – Trx | CL7 | 20-Amino Acid Linker | SUMO | Gene of Interest | His8 (This is the same as option 6, with the exception of His 8)
If XhoI is used in place of SpeI, the His8 tag is no longer in sequence.
The Bsu361/SpeI insertion scheme maintains the Gene of Interest’s wildtype sequence, without adding any extra residues. The N-terminus of the Gene of Interest must include the following to complete the SUMO C-terminal sequence (for options 3 and 6):


EXPIRATION | 6 months from receipt when stored as directed
Plasmid Sequences are available on the Plasmid Guide Page
Use of TriAltus products falls under our limited license agreement and is for non-profit research use only. For commercial use, please contact us for licensing.