TriAltus' highly pure, tag-free Cas9 protein is used for genome editing with CRISPR technology. Recombinant Streptococcus pyogenes Cas9 (wild-type) protein is expressed in Escherichia coli and features two nuclear localization signals (NLS), one at the N-terminus and another at the C-terminus. Cas9 protein combines with guide RNA (gRNA) to form a stable ribonucleoprotein (RNP) complex that enables efficient and precise genome editing in cells.
SPECIFICATIONS
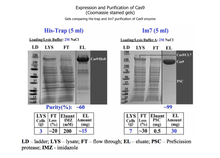
PURITY | ~99% (SDS-PAGE gel analysis)
ENZYME SOURCE | Streptococcus pyogenes Cas9 is expressed in E. coli with two (N-terminal and C-terminal) nuclear localization signals (NLS), and is purified with our CL7 tag technology.
ENDOTOXIN | <0.3 EU/ug by rFC method
STORAGE/SHIPPING CONCENTRATION | 5 mg/mL
STORAGE BUFFER | 10 mM Tris-Cl pH 8.0, 250 mM NaCl, 50% glycerol
RECOMMENDED STORAGE CONDITIONS | -80°C
EXPIRATION | 6 months from receipt when stored as directed. Avoid repeated freeze/thaw cycles.

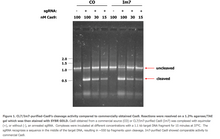
Figure 1. CL7/Im7-purified-Cas9’s cleavage activity compared to commercially obtained Cas9. Reactions were resolved on a 1.2% agarose/TAE gel which was then stained with SYBR GOLD. Cas9 obtained from a commercial source (CO) or CL7/Im7-purified Cas9 (Im7) was complexed with equimolar (+), or without (-), an annealed sgRNA. Complexes were incubated at different concentrations with a 1.1 kb target DNA fragment for 15 minutes at 37°C. The sgRNA recognizes a sequence in the middle of the target DNA, resulting in ~550 bp fragments upon cleavage. Im7-purified Cas9 showed comparable activity to commercial Cas9.

Figure 2. Activities of CL7/Im7-purified Cas9 and 4-step purified Cas9 are the same in cells.
(A) Phase contrast microscopy of human sickle cell iPSCs (induced pluripotent stem cells) after electroporation with Lonza nucleofector 2b and varying concentrations of RNP using Cas9WT from either the 4-step purification (top row) or 1-step purification protocols (bottom row).
(B) Table of viabilities and modifications from the electroporation experiments shown in (A). RNPs made with Cas9 purified using the two protocols were tested using a sgRNA designed to correct a 1-bp error in hemoglobin that causes sickle cell disease. Modification rates were evaluated by digital PCR. The proteins resulted in the same viability and modification rates, indicating equivalent efficiency in cells. These tests were performed by Tim Townes and Lei Ding of the University of Alabama at Birmingham (UAB).
SKU |
100 µg: 40-1320
500 µg: 40-1325
Customize concentration and scale to meet your specific research needs. Please Inquire.
Use of TriAltus products falls under our limited license agreement and is for non-profit research use only. For commercial use, please contact us for licensing.